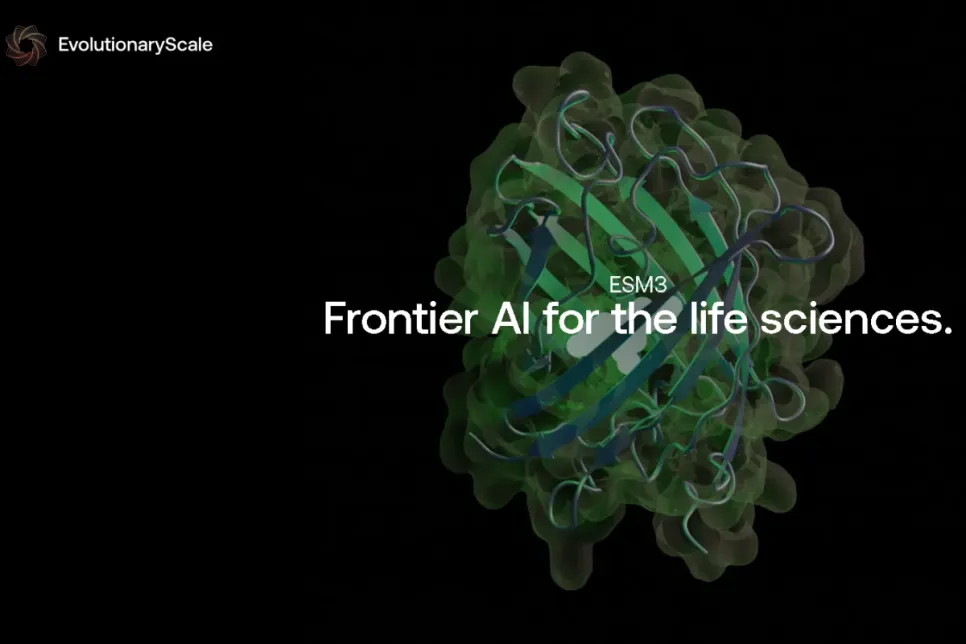
EvolutionaryScale, a frontier AI research lab for biology, launched with ESM3, an AI model capable of generating novel proteins. ESM3 generated a new Green Fluorescent Protein (GFP), a process that would take 500 million years of evolution to occur naturally. This milestone generative AI model allows interactive prompting to create proteins, empowering scientists to advance applications from drug discovery, and materials science, to carbon capture.
The founding team at EvolutionaryScale and behind ESM3 are pioneers in applying AI to biology, building what is widely considered to be the first transformer language model for proteins ESM1. The ESM models have empowered groundbreaking scientific research, including a breakthrough in protein folding that helped reveal the structures of hundreds of millions of metagenomic proteins; the models have been used by scientists across the world to model and understand proteins.
ESM3 was trained with 1 trillion teraflops – more compute than any other known model in biology – on a dataset of 2.78 billion proteins across the Earth’s natural diversity. It is the first generative model for biology that simultaneously reasons over the sequence, structure, and function of proteins. This enables scientists to understand and create new proteins, making biology programmable.
“ESM3 takes a step toward a future of biology where AI is a tool to engineer from first principles, the way we engineer structures, machines, and microchips, and write computer programs,” said EvolutionaryScale co-founder and chief scientist, Alexander Rives. “We’ve been working on this for a long time, and we’re excited to share it with the scientific community and see what they do with it.” With this capability, the model has the potential to accelerate discovery across a broad range of applications, ranging from the development of new cancer treatments to the creation of proteins that could help capture carbon.
Prompted through a chain of thought to reason over possible sequences and structures of GFP, ESM3 stepped across 500 million years of evolution to create a new fluorescent protein. GFP is one of the most beautiful and unique proteins in nature, responsible for the glowing of jellyfish and the vivid fluorescent colors of coral. It is the only protein that emits light, and the biological mechanism for this is unique – it is a protein that transforms itself forming a light-emitting chromophore out of its own atoms.
GFP has become an important tool in molecular biology, helping scientists to see molecules inside cells. The mechanism that powers this phenomenon is incredibly complex, and generating a variant this distant by computational or experimental laboratory techniques has not been scientifically documented. New fluorescent proteins that are distant from known ones have only been found through the discovery of new GFPs in the natural world. The analysis suggests that under natural evolution it could take more than 500 million years for a protein this different to evolve.
ESM3’s success in generating a new GFP underscores the model’s potential for advancements in biological research and life sciences. EvolutionaryScale will be opening an API for closed beta today and code and weights are available for a small open version of ESM3 for non-commercial use. EvolutionaryScale is also collaborating with AWS and NVIDIA to accelerate applications from drug discovery to synthetic biology with AI.
EvolutionaryScale also announced a seed round of more than $142 million, led by Nat Friedman and Daniel Gross, and Lux Capital, with participation from Amazon, NVentures (NVIDIA’s venture capital arm), and angel investors. Funding will be used to further expand the capabilities of its models.